

Biometric Research Branch home page. STAR: Biogene - Home. Pathguide: the pathway resource list. SOURCE Search. SOURCE is a unification tool which dynamically collects and compiles data from many scientific databases, and thereby attempts to encapsulate the genetics and molecular biology of genes from the genomes of Homo sapiens, Mus musculus, Rattus norvegicus into easy to navigate GeneReports. The mission of SOURCE is to provide a unique scientific resource that pools publicly available data commonly sought after for any clone, GenBank accession number, or gene. Database of Protein and Genetic Interactions. Taverna - open source and domain independent Workflow Management System. BioCatalogue Services. The R Project for Statistical Computing.
Statconn. Bioconductor - Home. 2.8 Software Packages. R Commander. John Fox and Milan Bouchet-Valat Please Read the Rcmdr Installation Notes (click on the image for a larger view) For more details, see my paper on the R Commander in the Journal of Statistical Software (which is somewhat out of date) and the introductory manual distributed with the package (accessible via the Help -> Introduction to the R Commander menu).

The R-Commander GUI consists of a window containing several menus, buttons, and information fields. (The menu tree, etc., are shown below.) The menus lead to simple dialog boxes, the general contents of which are more or less obvious from the names of the menu items. By default, commands generated via the dialogs are posted to the output window, along with printed output, and to the script window. Commands access a current or active data set (data frame). In addition to standard packages, the R Commander uses functions in a number of other packages. I'm making the GUI available as the Rcmdr package. R Tutorial. 321a Boyd Graduate Studies University of Georgia Athens, Georgia 30602 Introductory Materials¶ These materials are designed to offer an introduction to the use of R.
It is not exhaustive, but is designed to just provide the basics. Introduction to R. RStudio - Home. Systems-Biology. BiologicalNetworks. The STRING database in 2011: functional interaction networks of proteins, globally integrated and scored. Proteins can form a variety of functional connections with each other, including stable complexes, metabolic pathways and a bewildering array of direct and indirect regulatory interactions.

These connections can be conceptualized as networks and the size and complex organization of these networks present a unique opportunity to view a given genome as something more than just a static collection of distinct genetic functions. Indeed, the ‘network view’ on a genome is increasingly being taken in many areas of applied biology: protein networks are used to increase the statistical power in human genetics (1,2), to aid in drug discovery (3,4), to close gaps in metabolic enzyme knowledge (5,6) and to predict phenotypes and gene functions (7,8), to name just a few examples.
STRING: functional protein association networks. Psicquic - Protemics Standard Initiative Common QUery InterfaCe. This project hosts the PSICQUIC Reference Implementation, an implementation of the PSICQUIC Specification.
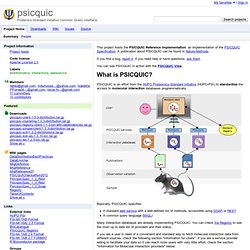
A publication about PSICQUIC can be found in Nature Methods. If you find a bug, report it. If you need help or have questions, ask them. You can see PSICQUIC in action with the PSICQUIC View. PSICQUIC is an effort from the HUPO Proteomics Standard Initiative (HUPO-PSI) to standardise the access to molecular interaction databases programmatically. Basically, PSICQUIC specifies: A standard web service with a well-defined list of methods, accessible using SOAP or REST. Many interaction databases are already implementing PSICQUIC. PSICQUIC View. This application allows you to query PSICQUIC services and browse their respective interactions.
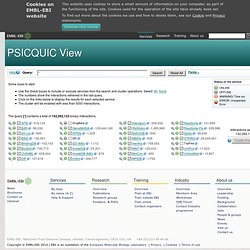
However these dataset can feature multiple evidence of two molecule interacting together, not only within one service but also across several. The same, redundant information may have been separately curated by multiple resources. Ingenuity Systems. Your Favorite Gene (YFG): gene specific search results for shRNAs, antibodies & cofactors. Discover how our website is making it easier for you to find information and products.
Search for products supported by peer-reviewed full-text papers that are integrated with curated scientific information. We have incorporated the gene content from Ingenuity (formerly known as Your Favorite Gene) within our search results, product information and papers. Now, finding information related to your gene is easier than ever. Find products, papers and gene information in a single search. Read peer-reviewed papers that elucidated the interactions you are interested in. Cytoscape: An Open Source Platform for Complex-Network Analysis and Visualization. Download NetAtlas software for free. NetAtlas: A Cytoscape plugin to examine signaling networks based on tissue gene expression.
Data_Sets - Cytoscape Wiki. Example Data Sets These data sets have been generously contributed by users.
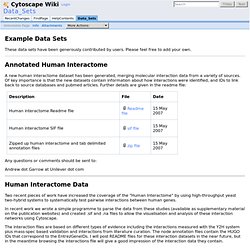
Please feel free to add your own. Presentations - Cytoscape Wiki. Cytoscape Web: an interactive web-based network browser. + Author Affiliations * To whom correspondence should be addressed.

Received May 3, 2010. Revision received July 1, 2010. Accepted July 20, 2010. BioTapestry. Systems Biology - BioChemWeb.org. PubMed home. SOURCE: a unified genomic resource of functional annotations, ontologies, and gene expression data. Nucleic Acids Research Oxford University Press Abstract The explosion in the number of functional genomic datasets generated with tools such as DNA microarrays has created a critical need for resources that facilitate the interpretation of large-scale biological data.

SOURCE is a web-based database that brings together information from a broad range of resources, and provides it in manner particularly useful for genome-scale analyses. SOURCE's GeneReports include aliases, chromosomal location, functional descriptions, GeneOntology annotations, gene expression data, and links to external databases. Molecular Systems Biology. Molecular Systems Biology. Abstract Subtelomeric chromatin is subject to evolutionarily conserved complex epigenetic regulation and is implicated in numerous aspects of cellular function including formation of heterochromatin, regulation of stress response pathways and control of lifespan.

Subtelomeric DNA is characterized by the presence of specific repeated segments that serve to propagate silencing or to protect chromosomal regions from spreading epigenetic control. In this study, analysis of genome‐wide chromatin immunoprecipitation and expression data, suggests that several yeast transcription factors regulate subtelomeric silencing in response to various environmental stimuli through conditional association with proto‐silencing regions called X elements.
Visual Overview. Molecular Systems Biology. Abstract Side effect similarities of drugs have recently been employed to predict new drug targets, and networks of side effects and targets have been used to better understand the mechanism of action of drugs. Here, we report a large‐scale analysis to systematically predict and characterize proteins that cause drug side effects. We integrated phenotypic data obtained during clinical trials with known drug–target relations to identify overrepresented protein–side effect combinations. PLoS Collections: Article collections published by the Public Library of Science. Reactome. Reactome Introduction User Series 1.