

Exome Sequencing vs RNA-Seq to Identify Coding Region Variants. Whole Genome Sequencing The advent of next-generation sequencing technologies has revolutionized the study of genetic variation in the human genome.
Whole-genome sequencing currently represents the most comprehensive strategy for variant detection genome-wide but is costly for large sample sizes, and variants detected in noncoding regions remain largely uninterpretable. Global Next Generation Sequencing (NGS) Market Worth $2,343 Million by 2016. From: Next Generation Sequencing (NGS) Market – Global Trends and Forecasts (2011-2016), marketsandmarkets.com, April 2012 The global NGS market was valued at $842.5 million in the year 2011, growing at a CAGR of 22.7% from 2012 to 2016.
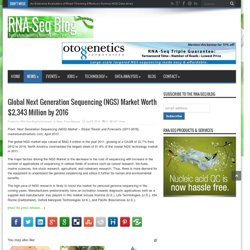
North America commanded the largest share of 51.8% of the overall NGS technology market in 2011. The major factors driving the NGS Market is the decrease in the cost of sequencing with increase in the number of applications of sequencing in various fields of science such as cancer research, bio-fuels, marine sciences, live stock research, agricultural, and veterinary research. Thus, there is more demand for the equipment to understand the genome sequencing and utilize it further for human and environmental benefits. What the ‘limits of DNA’ story reveals about the challenges of science journalism in the ‘big data’ age. As a science journalist, I sympathize with book reviewers who wrestle with the question of whether to write negative reviews.
It seems a waste of time to write about a dog of a book when there are so many other worthy ones; but readers deserve to know if Oprah is touting a real stinker. On 2 April, Science Translational Medicine published a study on DNA’s shortcomings in predicting disease. My editors and I had decided not to cover the study last week after we saw it in the journal’s embargoed press packet, because my sources offered heavy critiques of its methods. But it was a tough choice: we knew the paper was bound to get a lot of other coverage, as it conveyed a provocative message, would be published in a prominent journal, and would be highlighted at a press conference at the well-attended annual meeting of the American Association for Cancer Research.
What the ‘limits of DNA’ story reveals about the challenges of science journalism in the ‘big data’ age. Seq of Bacteria. A Team of researchers at the Broad Institute set out to establish a robust and scalable RNA-seq process applicable to cultured bacteria as well as to complex community transcriptomes.
They determined that an effective process should: a) reduce rRNA sequences to very low levels b) accurately maintain relative representation of transcript sequences. RNA-Seq reveals complexity of the Arabidopsis transcriptome. Researchers at the Medical University of Vienna have conducted a genome-wide study analyzing alternative splicing (AS) in Arabidopsis.
AS is a key regulatory mechanism that contributes to transcriptome and proteome diversity. They performed high-throughput sequencing of a normalized cDNA library which resulted in a high coverage transcriptome map of Arabidopsis. Optimizing a Massive Parallel Sequencing Workflow for Quantitative miRNA Expression Analysis. Massive Parallel Sequencing methods (MPS) can extend and improve the knowledge obtained by conventional microarray technology, both for mRNAs and short non-coding RNAs, e.g. miRNAs.
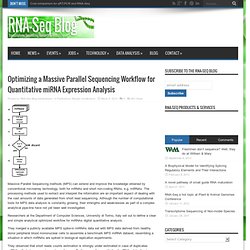
The processing methods used to extract and interpret the information are an important aspect of dealing with the vast amounts of data generated from short read sequencing. Although the number of computational tools for MPS data analysis is constantly growing, their strengths and weaknesses as part of a complex analytical pipe-line have not yet been well investigated.
Insights into Chinese prostate cancer with RNA-seq. Prostate cancer remains a leading cause of cancer morbidity and mortality in men, accounting for approximately one million new cases and 260 000 deaths per year worldwide.
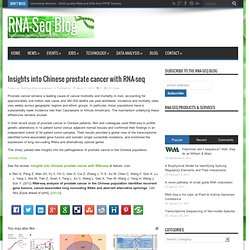
Incidence and mortality rates vary widely across geographic regions and ethnic groups. In particular, Asian populations have a substantially lower incidence rate than Caucasians or African Americans. Webinar – Analyzing the Cancer Transcriptome – Illuminating the Dysfunctional Cancer Genome. Presented by GEN Broadcast Date: Wednesday, April 18, 2012Time: 1 pm ET, 10 am PT The RNA seq method provides a comprehensive amount of information on the cancer transcriptome: highly sensitive and accurate gene-expression levels, alternative splice forms, novel fusion transcripts, and mutational changes that have a direct impact on protein function.
In this webinar, presenters will review transformative discoveries from RNA-seq, and describe a highly effective approach for isolation and characterization of circulating tumor cells using RNA-seq at single cell resolution. Parameters that affect quantification of expressed nucleotide sequence polymorphisms (SNVs) will also be discussed, as will the development and implementation of tools for the identification of fusion transcripts. Finally, presenters will also talk about development of a potential RNA-seq based test for cancer. Seq of Bacteria. Seq at AACR 2012. The annual meeting of the American Association of Cancer Research (AACR) will be held next week in Chicago.
RNA-Seq is one of many new technologies enabling significant discoveries in the areas of cancer research, diagnosis and treatment. Here is just a partial list of some of the presentations related to RNA-Seq that you can attend at the meeting next week. RNA-Seq Data Analysis Workshop. Optimizing a Massive Parallel Sequencing Workflow for Quantitative miRNA Expression Analysis. Protocol – Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. RNA-Seq reveals complexity of the Arabidopsis transcriptome. Seq Workshop. RNA Advances Reshape Prevailing Wisdom. From Genetic Engineering News Expanding on a concept originally coined by Walter Gilbert in 1986, Thomas Cech recently described two RNA worlds—a hypothetical, primordial world in which the same molecule combined informational and catalytic properties, and a contemporary world, forged by a spectrum of RNA-centered activities.
While during the early days, following the discovery of DNA, RNA was thought to be the less interesting molecule, this view has dramatically changed in recent years. Advances in RNA biology are reshaping concepts that have been prevailing for a long time. For example, in the field of developmental biology, decisions were traditionally envisioned mostly in terms of the progressive expression of different combinations of transcription factors that are induced by specific combinations and concentrations of growth differentiation factors. (read more…) Sugar Beet Transcriptome. Sugar beet (Beta vulgaris sp. vulgaris) crops account for about 30% of world sugar. Sugar yield is compromised by reproductive growth hence crops must remain vegetative until harvest.
There is no sugar beet reference genome, or public expression array platforms. RNA-Seq has now enabled the generation of the first reference transcriptome for sugar beet and the study of global transcriptional responses in the shoot apex to vernalization and GA treatment, without the need for a reference genome or established array platforms. Comprehensive bioinformatic analysis identified transcriptional programmes associated with different sugar beet genotypes as well as biological treatments; thus providing important new opportunities for basic scientists and sugar beet breeders.
Transcriptome-scale identification of agronomically important traits as used in this study should be widely applicable to all crop plants where genomic resources are limiting. Tutorial on RNA-seq mapping and data analysis. Maize (Zea mays L.) Genome Diversity as Revealed by RNA-Sequencing. Maize (Zea mays L.) Genome Diversity as Revealed by RNA-Sequencing. Seq Tutorial from the iPlant Collaborative. Insights into Chinese prostate cancer with RNA-seq. RNA-Seq Blog. Transcriptome Sequencing Identifies Copper Acquisition Genes.
If you were wondering about which genes are involved in the copper dependence of iron homeostasis in Arabidopsis, here’s your answer! Turns out its: SQUAMOSA PROMOTER BINDING PROTEIN-LIKE7 (SPL7), FERRIC REDUCTASE OXIDASE5 (FRO5) and FERRIC REDUCTASE OXIDASE4 (FRO4). Yep, researchers at Ruhr University, Germany discovered this with transcriptome sequencing. RNA-Seq sheds light on dysregulation of vascular endothelial cells by thrombin. The dysregulation of vascular endothelial cells by thrombin has been implicated in the development of a number of pathologic disorders such as inflammatory conditions, cancer, diabetes, coronary heart disease. However, transcriptional regulation of vascular endothelial cells by thrombin is not completely understood. Researchers at the University of Missouri, used Illumina RNA-Seq to profile the transcriptome in human pulmonary microvascular endothelial cells (HMVEC-L) treated with thrombin for 6 h to gain insight into thrombin’s direct effects on the endothelial function.
RNA-Seq to Measure Protein Abundance. Within the proteomics community there is a substantial interest in development of novel label-free quantitative proteomic strategies. One strategy is to take advantage of an increasing number of studies involving integrative analysis of gene and protein expression data that are based on new technologies such as next-generation transcriptome sequencing (RNA-Seq) and highly sensitive mass spectrometry (MS) instrumentation. Thus, it becomes interesting to revisit the correlative analysis of gene and protein expression data using more recently generated datasets to determine if gene expression data can be used as an indirect benchmark for such protein-level comparisons. A team of researchers from the University of Michigan and the Chinese Academy of Sciences used publicly available mouse data to perform a joint analysis of genomic and proteomic data obtained on the same organism. Groundbreaking Research – mRNA sequencing reveals transcriptomic landscape of breast cancer.
A team led by researchers at the George Washington University has published a study that is the first of its kind to use mRNA sequencing to look at the expression of genome, at a unprecedented resolution at the current time, in three types of breast cancer. The study titled, “Transcriptomic landscape of breast cancer through mRNA sequencing,” is published in the Feb. 14 edition of the journal, Scientific Reports, a new open access Nature journal for large volume data.
Breast cancer is a heterogeneous disease with a poorly defined genetic landscape, which poses a major challenge in diagnosis and treatment. By massively parallel mRNA sequencing, the team obtained 1.2 billion reads from 17 individual human tissues belonging to TNBC, Non-TNBC, and HER2-positive breast cancers and defined their comprehensive digital transcriptome for the first time.
(read more… ) RNA-Seq Blog. Prognosys Launches RNA-Seq Study-Level Statistical Module for the Voila! Analysis Platform. RNA-Seq Workshops. RNA-Seq sheds light on dysregulation of vascular endothelial cells by thrombin. Transcriptome Analysis of the Model Protozoan – Tetrahymena thermophila, Using Deep RNA Sequencing. Modeling RNA degradation for RNA-Seq. Shareholders sue Illumina over Roche bid. RNA-Seq Atlas – a web-based repository of RNA-Seq gene expression profiles and query tools. Expression Analysis Announces RNA-SEQ Grant Program. De Novo Assembly of the Manila Clam Ruditapes philippinarum Transcriptome. Groundbreaking Research – mRNA sequencing reveals transcriptomic landscape of breast cancer. Leveraging in Statistical Analysis of RNA-Seq. Life Technologies weighs in on Roche – Illumina Deal. The Pseudomonas aeruginosa Transcriptome.