

mRNA - Accepts Sequence of Codons (mRNA) and Translates this Sequence into Protein. My Biosoftware. ELL 3.2.3 - Platform for Generic Algorithms. My Biosoftware. CoRAL 1.1.1 - Classification of RNAs by Analysis of Length. GOseq 1.12.0 - Performing Gene Ontology (GO) based tests on RNA-seq data. Subread 1.3.5-p5 / RSubread 1.10.5 - Processing Next-gen Sequencing data. ReverseComplement 1.8 - Reverse Complement of DNA and RNA sequences. The RNomics-structbio #Paper. The RNomics-RNA #Paper. My Biosoftware. BioFace 0.02 - Simplified tool for Genetic Analyzes. Rna-star 2.2.0 - Spliced Transcripts Alignment to a Reference. DIAL - 3-Dimensional RNA Structural Alignment and Motif Detection.
Rnaifold - RNA Inverse Folding and Molecular Design. RNAmapper v1 - Use of RNA-Seq data to Map and Identify Mutations in Zebrafish. ReadSpy - Assessment of Uniformity in RNA-Seq reads. Rnapagenumber - Compute Optimal "page number" of an RNA Structure. RNAbor - Compute Structural Neighbors of an RNA Secondary Structure. GP 0.26 / Arka 0.11 - Sequence Manipulation tools. Get_size_NN v1 - Neutral Network Size Estimation. CURE 1.1 - Cytidine-to-Uridine RNA Editing Predictor. Starfold - Predict RNA Secondary Structures. FindtRNA 1.0 - tRNA Prediction tool. MapMi 1.5.0-b01 - Automated Mapping of miRNA loci. fRNAkenstein 20120710 - Python program for Multiple Target Inverse RNA Folding.
DNA to RNA 1.0 - Converts DNA code to mRNA code. DNA/RNA/Protein and General Molecular Weight Calculator 1.0. RNAmotifs 3 - Predict a motif for your own set of RNAs. Polyadq - Detection of Human Polyadenylation Signals. PIPAx 2.1.1 - Bioinformatics Analysis and Management of NGS data. Tarsier - Testing and Analysing RNA gene Software, Including Evolutionary Relationships. TaLasso - Quantification of miRNA-mRNA interactions. The TaLasso (miRNA-Target LASSO) web site is an easy tool where only few steps are needed to get the interaction scoring results provided by TaLasso algorithm Alberto Pascual Montano Windows/Linux/MacOsXMatlab / R Package TaLasso Citation PLoS One. 2012;7(2):e30766.
SpliceGrapher 0.0.5 - Creat Splice Graphs from RNA-Seq data. RNAAnalyzer - Identify Regulatory RNA Elements. RNAAnalyzer is an integrated tool-box to identify regulatory RNA elements.
The RNA analyzer collects general and specific information on any submitted RNA sequence or batch of sequences in FASTA format. It determines and rapidly scans the different regions of an RNA (including 5′ UTR, CDS, 3′ UTR in mRNA) and screens for specific RNA signals (in each of these regions, e.g. polyA-site, AU rich region etc. in 3′ UTR). It runs a fast folding RNA routine to provide an overview of the RNA fold. Furthermore it analyzes structure content, fold energy and stem loops. the Department of Bioinformatics at the University of Würzburg Web Browser Citation.
WAR - Webserver for Aligning structural RNAs. EteRNA - RNA Game Lets Players Help Find a Biological Prize. By playing EteRNA, you will participate in creating the first large-scale library of synthetic RNA designs.
Your efforts will help reveal new principles for designing RNA-based switches and nanomachines — new systems for seeking and eventually controlling living cells and disease-causing viruses. By interacting with thousands of players and learning from real experimental feedback, you will be pioneering a completely new way to do science. the Das Lab Web BrowserFlash Citation. Caryoscope 0.4.0 - View Gene Expression Data in Genome. Caryoscope is an application for viewing gene expression data in a whole-genome context.
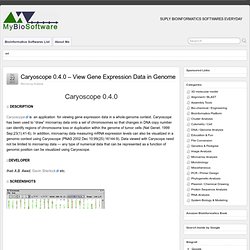
Caryoscope has been used to “draw” microarray data onto a set of chromosomes so that changes in DNA copy number can identify regions of chromosome loss or duplication within the genome of tumor cells (Nat Genet. 1999 Sep;23(1):41-6). In addition, microarray data measuring mRNA expression levels can also be visualized in a genomic context using Caryoscope (PNAS 2002 Dec 10;99(25):16144-9). Data viewed with Caryscope need not be limited to microarray data — any type of numerical data that can be represented as a function of genomic position can be visualized using Caryoscope. PARE - Compare Protein Abundance and mRNA Expression data. PARE (Protein Abundance and mRNA Expression) is a tool for comparing protein abundance and mRNA expression data.
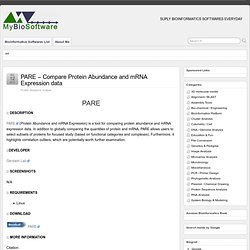
In addition to globally comparing the quantities of protein and mRNA, PARE allows users to select subsets of proteins for focused study (based on functional categories and complexes). Furthermore, it highlights correlation outliers, which are potentially worth further examination. Gerstein Lab Linux. Riboswitch finder - Identification of Riboswitch RNAs. Riboswitch finder is a dedicated RNA motif search program and web server to identify RNA riboswitches.
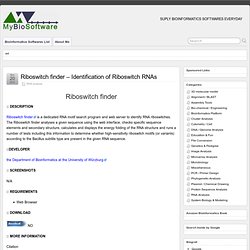
The Riboswitch finder analyses a given sequence using the web interface, checks specific sequence elements and secondary structure, calculates and displays the energy folding of the RNA structure and runs a number of tests including this information to determine whether high-sensitivity riboswitch motifs (or variants) according to the Bacillus subtilis type are present in the given RNA sequence. the Department of Bioinformatics at the University of Würzburg Web Browser Citation Riboswitch finder–a tool for identification of riboswitch RNAs. BRAGI 20091106 - A Protein Visualization and Modeling Program. BRAGI is a well-established package for viewing and modeling of three-dimensional (3D) structures of biological macromolecules.BRAGI enables you to view and explore the three-dimensional (3D) structure of any macromolecule.
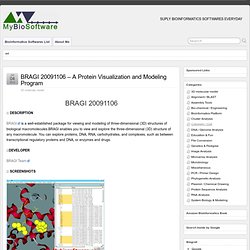
You can explore proteins, DNA, RNA, carbohydrates, and complexes, such as between transcriptional regulatory proteins and DNA, or enzymes and drugs. BRAGI Team Windows / Linux Citation BRAGI: linking and visualization of database information in a 3D viewer and modeling tool. PhenoFam - Gene Set Enrichment Analysis in the Protein Domain Context. PhenoFam performs gene set enrichment analysis by employing structural and functional information on families of protein domains as annotation terms.
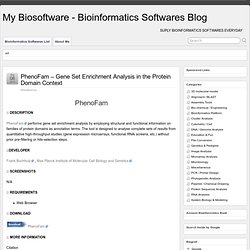
The tool is designed to analyse complete sets of results from quantitative high-throughput studies (gene expression microarrays, functional RNAi screens, etc.) without prior pre-filtering or hits-selection steps. Frank Buchholz , Max Planck Institute of Molecular Cell Biology and Genetics. Web Browser PhenoFam Citation. RLooM - RNA Loop Modeling. RLooM is a web application for homology-based modeling of RNA loops utilizing template structures extracted from the PDB.
RLooM allows the insertion and replacement of loop structures of a desired sequence into an existing RNA structure. Furthermore, a comprehensive database of loops in RNA structures can be accessed through the web interface. Max Planck Institute for Molecular Plant Physiology Web Browser. IQSeq 1.0 - Isoform Quantification with RNA-seq data. IQSeq is a tool for isoform quantification with RNA-seq data.
Given isoform annotation and alignment of RNA-seq reads, it will use an EM algorithm to infer the most probable expression level for each isoform of a gene. Gerstein Lab Linux IQSeq Citation: Jiang Du, Jing Leng, Lukas Habegger, Andrea Sboner, Drew McDermott, Mark Gerstein. FRASS 1.0 - RNA Structure Comparison. The FRASS web-server represents a RNA chain with its Gauss integrals and allows one to compare structures of RNA chains and to find similar entries in a database derived from the Protein Data Bank. We observed that FRASS scores correlate well with the ARTS and LaJolla similarity scores. Moreover, the-web server can also reproduce satisfactorily the DARTS classification of RNA 3D structures and the classification of the SCOR functions that was obtained by the SARA method. Silvio Tosatto LinuxC++ Compiler Citation: RseqFlow - RNA-seq Data Analysis Workflow.
RseqFlow is a workflow which can implement a set of RNA-Seq analysis, such as performing quality control (QC) for sequencing data, generating signal tracks of mapped reads, calculating expression levels, and identifying differentially expressed genes. The workflow allows users to express multi-step RNA-Seq analysis by just providing sequencing RNA-Seq datasets and typing a few commands in an easy-to-use environment. Ting Chen Linux/ Windows / MacOsXVirtual Machine. WEAV 0.2 - de novo Assembly program for both Genome and RNA. RobiNA 1.2.3 - Open Source Microarray and RNA-Seq Processing. RobiNA is an integrated solution that consolidates all steps of RNA-Seq-based differential gene-expression analysis in one user-friendly cross-platform application featuring a rich graphical user interface.
RobiNA accepts raw FastQ files, SAM/BAM alignment files and counts tables as input. It supports quality checking, flexible filtering and statistical analysis of differential gene expression based on state-of-the art biostatistical methods developed in the R/Bioconductor projects. In-line help and a step-by-step manual guide users through the analysis. Max Planck Institute for Molecular Plant Physiology, Golm, Germany. Windows/Linux/MacOsXJava. CROP 1.33 - Clustering 16S rRNA For OTU Prediction. CROP is a clustering tool designed mainly for Metagenomics studies, which clusters 16S rRNA sequences into Operational Taxonomic Units (OTU). Ting Chen LinuxGSLC++ Compiler Citation: Xiaolin Hao; Rui Jiang; Ting ChenClustering 16S rRNA for OTU prediction: a method of unsupervised Bayesian clustering Bioinformatics 2011; doi: 10.1093/bioinformatics/btq725.
CleaveLand 3.0.1 - Find Cleaved Small RNA Targets. CleaveLand is a command-line executed pipeline for finding cleaved small RNA targets using degradome data (also known as PARE [parallel analysis of RNA ends] and GMUCT [genome-wide mapping of uncapped transcripts]). Provided with a set of degradome data, a list of small RNA queries, a reference transcriptome/mRNA set, CleaveLand outputs potentially cleaved small RNA targets along with other supporting information. Axtell Lab @ Penn State CleaveLand Citation: NePhe 1.0 - Network RNAi Phenotype Score. BBSeq 1.0 - Analysis of RNA Sequence Count Data.
BBSeq (Beta-Binomial modeling of the overdispersion of the RNA-seq count data)is used to identify differential expression in high-throughput count data, such as RNAseq count data which is derived from next-generation sequencing machines. Our modeling design is very flexible. It can not only solve the data with multiple comparisons, but also can find the affect from other covariates, such as age and other counfounding variables. NOISeq - Differential Expression in RNA-seq. NOISeq is a non-parametric approach for the identification of differentially expressed genes from count data. NOISeq empirically models the noise distribution of count changes by contrasting fold-change differences (M) and absolute expression differences (D) for all the features in samples within the same condition.
GENE-E 1.0 - Exploration of data sets derived from RNAi. GENE-E (Gene Experiment) is a Java desktop application developed to allow the rapid visual exploration of data sets derived from RNAi and chemical screens. GENE-E’s user interface is based on an interactive heat map that allows users to easily highlight and drill down to regions of interest. rSeq 0.0.7 - RNA-Seq Analyzer. SpliceMap 3.3.5.2 - Splice Junction Discovery Using RNA-Seq. SpliceMap is a de novo splice junction discovery and alignment tool. CONSAN 1.2 - Pairwise Structural RNA Alignment. QRNA 2.03d - Prototype Noncoding RNA Genefinder. PKNOTS 1.07 - RNA Pseudoknot Prediction. RNAML 1.1.2 - Syntax for Exchanging RNA Information.
RNAML is a standard syntax for exchanging RNA information.This RNAML syntax allows for the storage and the exchange of information about RNA sequence and secondary and tertiary structures. The syntax permits the description of higher level information about the data including, but not restricted to, base pairs, base triples, and pseudoknots Laboratoire de Bioinformatique Théorique. GENE-E 1.0 - Exploration of data sets derived from RNAi. Trinity 20111126 - RNA-Seq De novo Assembly. Trinity represents a novel method for the efficient and robust de novo reconstruction of transcriptomes from RNA-Seq data. Trinity combines three independent software modules: Inchworm, Chrysalis, and Butterfly, applied sequentially to process large volumes of RNA-Seq reads.
Trinity partitions the sequence data into many individual de Bruijn graphs, each representing the transcriptional complexity at at a given gene or locus, and then processes each graph independently to extract full-length splicing isoforms and to tease apart transcripts derived from paralogous genes. The Broad Institute, Cambridge, MA. PALMapper 0.4 - Spliced Alignments of RNA-Seq Reads. BLISS 0.7 - Identify Batch Effects in RNA Expression Data. Qcalculator 1.0 - Calculate Relative mRNA Gene Expression. RALEE 0.61 - RNA ALignment Editor in Emacs.
RALEE is a major mode for the Emacs text editor. It provides functionality to aid the viewing and editing of multiple sequence alignments of structured RNAs. It is extremely immature, but is already proving useful to Rfam people. SGJlab Citation: SARSE 1.37 - RNA Sequence Editor. SARSE (semiautomated RNA sequence editor) is a graphical sequence editor for working with structural alignments of RNA. RNAdbtool - Update and Cleanup of Structural RNA Databases. LocARNA 1.6.2 - Global and Local Alignment of RNA. IntaRNA 1.2.5 - RNA-RNA Interaction Prediction. INFO-RNA 2.1.2 - A Server for Inverse Folding of RNA. LocalFold 1.0 - Local Folding of RNA. MARNA 100729 – Server for Multiple Alignment of RNAs. MEMERIS 1.0 - MEME in Rna's Including Secondary Structures. PiRNA 3.3 - RNA Interaction Search Engine. ARTS - Alignment of RNA Tertiary Structures. Darn 1.0 - Non-protein-coding RNA Detection. NcFANs - non-coding RNA Function Annotation Server. Erpin 5.5.4 - RNA Motif Search program.
RADAR 2.0 - RNA Data Analysis and Research.