

Autism study reveals a 'DNA tag' (methylation) amenable to treatment. A new discovery raises hope that autism may be more easily diagnosed and that its effects may be more reversible than previously thought.

In a new study appearing online in The FASEB Journal, scientists have identified a way to detect the disorder using blood and have discovered that drugs which affect the methylation state ("DNA tagging") of genes could reverse autism's effects. This type of drug is already being used in some cancer treatments.
"As the mother of a now 22-year-old son with an autism spectrum disorder, I hope that our studies as well as those of others, will lead to therapies that are designed to address specific deficiencies that are caused by autism, thus improving the lives of affected individuals," said Valerie W. Autism risk gene linked to differences in brain structure. Healthy individuals who carry a gene variation linked to an increased risk of autism have structural differences in their brains that may help explain how the gene affects brain function and increases vulnerability for autism.

The results of this innovative brain imaging study are described in an article in the groundbreaking neuroscience journal Brain Connectivity, a bimonthly peer-reviewed publication from Mary Ann Liebert, Inc. The article is available free online at the Brain Connectivity website. Genetic component of autism spectrum disorders may be moderate compared to environment, twin study suggests. After evaluating twin pairs in which at least one child has autism or autism spectrum disorder (ASD), researchers suggest that the shared environment may play a more substantial role in development of the condition than shared genes do, according to a report published Online First by Archives of General Psychiatry, one of the JAMA/Archives journals.

Current estimates suggest that 40 of every 10,000 children have autism, and prevalence rates for ASDs are about 1 percent, according to background information in the article. Studies of siblings have found a concordance rate (the likelihood that if one child has the disorder, others will as well) of up to 14 percent. The authors note that in previous studies of twins, concordance rates for autism were relatively high for identical (monozygotic) twins, but nonexistent for fraternal (dizygotic) twins.
Epigenetic changes shed light on biological mechanism of autism. Scientists from King's College London have identified patterns of epigenetic changes involved in autism spectrum disorder (ASD) by studying genetically identical twins who differ in autism traits.

The study, published in Molecular Psychiatry, is the largest of its kind and may shed light on the biological mechanism by which environmental influences regulate the activity of certain genes and in turn contribute to the development of ASD and related behaviour traits. ASD affects approximately 1 in 100 people in the UK and involves a spectrum of disorders which manifest themselves differently in different people. People with ASD have varying levels of impairment across three common areas: deficits in social interactions and understanding, repetitive behaviour and interests, and impairments in language and communication development. The researchers studied an epigenetic mechanism called DNA methylation.
New regulatory autism gene discovered. Mutations found in individuals with autism interfere with endocannabinoid signaling in the brain. Mutations found in individuals with autism block the action of molecules made by the brain that act on the same receptors that marijuana's active chemical acts on, according to new research reported online April 11 in the Cell Press journal Neuron.

The findings implicate specific molecules, called endocannabinoids, in the development of some autism cases and point to potential treatment strategies. "Endocannabinoids are molecules that are critical regulators of normal neuronal activity and are important for many brain functions," says first author Dr. Csaba Földy, of Stanford University Medical School. Neuromodulation. Neuromodulation is the physiological process by which a given neuron uses one or more neurotransmitters to regulate diverse populations of neurons. This is in contrast to classical synaptic transmission, in which one presynaptic neuron directly influences a single postsynaptic partner. Neuromodulators secreted by a small group of neurons diffuse through large areas of the nervous system, affecting multiple neurons.
Examples of neuromodulators include dopamine, serotonin, acetylcholine, histamine and others. Neuromodulation can be conceptualized as a neurotransmitter that is not reabsorbed by the pre-synaptic neuron or broken down into a metabolite. Evolution's gift may also be at the root of a form of autism. A recently evolved pattern of gene activity in the language and decision-making centers of the human brain is missing in a disorder associated with autism and learning disabilities, a new study by Yale University researchers shows.

"This is the cost of being human," said Nenad Sestan, associate professor of neurobiology, researcher at Yale's Kavli Institute for Neuroscience, and senior author of the paper. "The same evolutionary mechanisms that may have gifted our species with amazing cognitive abilities have also made us more susceptible to psychiatric disorders such as autism. " FMR1. FMR1 (fragile X mental retardation 1) is a human gene[1] that codes for a protein called fragile X mental retardation protein, or FMRP.[2] This protein, most commonly found in the brain, is essential for normal cognitive development and female reproductive function.
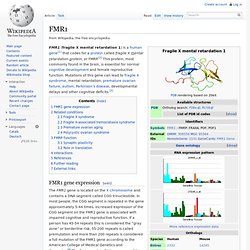
Mutations of this gene can lead to fragile X syndrome, mental retardation, premature ovarian failure, autism, Parkinson's disease, developmental delays and other cognitive deficits.[3] FMR1 gene expression[edit] The FMR1 gene is located on the X chromosome and contains a DNA segment called CGG trinucleotide. In most people, the CGG segment is repeated in the gene approximately 5-44 times. Increased expression of the CGG segment on the FMR1 gene is associated with impaired cognitive and reproductive function. The FMR1 gene can be found on the long (q) arm of the X chromosome at position 27.3, from base pair 146,699,054 to base pair 146,738,156.
Related conditions[edit] Fragile X syndrome[edit] Premature ovarian aging[edit]