

Roche vs Illumina – Round 2. Looking at micro could mend broken hearts. Ribonucleic acid (RNA) could hold the key to treating heart disease. The breakthrough could lead to the development of targeted therapeutic treatments for heart disease. Professor Thomas Preiss and Dr Jennifer Clancy and their team commenced the research at Sydney’s Victor Chang Cardiac Research Institute in 2008 and completed it at The John Curtin School of Medical Research at ANU. Last Month’s RNA-Seq Industry Press. New methods for quantifying differential expression with RNA-Seq. Rainbow Trout Transcriptome. Researchers at the USDA have generated and characterized a reference transcriptome for rainbow trout that represents multiple tissues responding to multiple stressors common to aquaculture production environments. This resource compliments existing public transcriptome data and will facilitate approaches aiming to evaluate gene expression associated with stress in this species.
Sequencing of a pooled normalized transcriptome library created from gill, brain, liver, spleen, kidney and muscle RNA of control and stressedfish produced 3,160,306 expressed sequence tags which were assembled and annotated. SNP discovery resulted in identification of ~58,000 putative single nucleotide polymorphisms including 24,479 which were predicted to fall within exons. C Sanchez CC, Weber GM, Gao G, Cleveland BM, Yao J, Rexroad CE. (2011) Generation of a reference transcriptome for evaluating rainbow trout responses to various stressors. BMC Genomics 12, 626. RNA-Seq a hot topic at Plant & Animal Genomes Conference. Sequencing Competition Heats Up! Illumina and Ion Torrent are two of the front runners in development of next-gen sequencing technology and it seems clear that both players believe there is only room for one top dog on this playground.
Back in the middle of 2011, Ion Torrent released an application note “The Ion PGM™ sequencer exhibits superior long-read accuracy” that, not only claimed better performance, but also seemed to stick it to Illumina by claiming they had accomplished in months what had taken Illumina years to accomplish. They also went as far as producing a humorous video, in the vein of Apple vs PC, attacking the Illumina MiSeq instrument. Illumina didn’t think it was funny and fired right back with its presentation, “Analysis of Inaccuracies in Ion Torrent’s Long Read Application Note” disputing the Ion Torrent claims and demonstrating better performance. Most recently, at the J.P. Transcriptomic analysis of Chinese bayberry (Myrica rubra) fruit development and ripening using RNA-Seq.
Chinese bayberry (Myrica rubra Sieb. and Zucc.) is an important subtropical fruit crop and an ideal species for fruit quality research due to the rapid and substantial changes that occur during development and ripening, including changes in fruit color and taste.
RNA-Seq generated 1.92 G raw data, which was then de novo assembled into 41,239 UniGenes with a mean length of 531 bp. Approximately 80% of the UniGenes (32,805) were annotated against public protein databases, and coding sequences (CDS) of 31,665 UniGenes were determined. Over 3,600 UniGenes were differentially expressed during fruit ripening, with 826 up-regulated and 1,407 down-regulated. GO comparisons between the UniGenes of these two types and interactive pathways (Ipath) analysis found that energy-related metabolism was enhanced, and catalytic activity was increased.
Bioinformatics Seminar – Using Finite Poisson Mixture Models for RNA-Seq Data Analysis and Transcript Expression Level Quantification. Bioinformatics Seminar – Using Finite Poisson Mixture Models for RNA-Seq Data Analysis and Transcript Expression Level Quantification. Effects of Sample Preparation on Transcriptome Sequencing Results. Seq of Cows Milk.
Cow milk is a complex bioactive fluid consumed by humans beyond infancy.
Even though the chemical and physical properties of cow milk are well characterized, very limited research has been done on characterizing the milk transcriptome. This study performs a comprehensive expression profiling of genes expressed in milk somatic cells of transition (day 15), peak (day 90) and late (day 250) lactation Holstein cows by RNA sequencing. Milk samples were collected from Holstein cows at 15, 90 and 250 days of lactation, and RNA was extracted from the pelleted milk cells. Gene expression analysis was conducted by Illumina RNA sequencing. Sequence reads were assembled and analyzed in CLC Genomics Workbench. Truly Digital RNA-Seq. Researchers at Harvard University have developed a method of truly digital RNA-Seq that does not rely on the counting of individual reads.
How is this possible you ask? Following reverse transcription, a large set of barcode sequences is added in excess, and nearly every cDNA molecule is uniquely labeled by random attachment of barcode sequences to both ends. After PCR, paired-end deep sequencing is performed to read the two barcodes and cDNA sequences. Rather than counting the number of reads, RNA abundance is measured based on the number of unique barcode sequences observed for a given cDNA sequence. The barcodes have been optimized to be unambiguously identifiable, even in the presence of multiple sequencing errors. The Challenge of RNA-Seq. The authors of a recent review1, make some key points about some of the challenges that are general to all RNA-seq experiments: RNA-Seq rules!
– “By comparison to these [other next-gen sequencing] applications, RNA-sequencing (RNA-seq) may be leading the pack in popularity because of its ability to characterize transcriptomes, to assess differential gene expression and to essentially challenge the continued use of microarray technology for studying transcription.” Transcriptome analysis of Crassostrea gigas with short read RNA-Seq. Researchers at the University of Washington used SOLiD sequencing to examine gene expression patterns in Pacific oyster (Crassostrea gigas) populations exposed to varying degrees of anthropogenic impact.
Novel transcripts were identified and RNA-Seq analysis revealed several hundred differentially expressed genes. To evaluate the effectiveness of restricting characterization solely to short-read sequences, mapping and RNA-Seq analysis were also performed using publicly available transcriptome sequence data as a scaffold. This study demonstrates that ultra-short read sequencing technologies can effectively generate novel transcriptome information, identify differentially expressed genes, and will be important for examining environmental physiology of non-model organisms.
Transcriptome analysis of Crassostrea gigas with short read RNA-Seq. Happy New Year! First Anolis carolinensis RNA-Seq data sets released. Simultaneous RNA-seq-based Transcript Inference and Quantification Using Mixed Integer Programming. Study Finds 454, Illumina Offer Similar Performance for Transcriptome Sequencing-Based SNP Discovery.
Seminar Tomorrow: RNA-Seq transcriptome analysis. RNA-Seq transcriptome analysis for gene identification, polymorphism detection and transcript profiling in two perennial ryegrass genotypes with divergent vernalization requirement By senior scientist Torben Asp, Department of Molecular Biology and Genetics, Aarhus University Thursday, 15 December 2011 at 10:00-11:00.
RNA-Seq Data Analysis Online Course. Seq Hands-on Workshop. RNA-Seq Satellite Workshop at ABRF 2012. Seq Blog on Pearltrees. RNA-Seq of the yeast Scheffersomyces stipitis. Xylose is the second most abundant lignocellulosic component besides glucose, but it cannot be fermented by the widely used ethanol-producing yeast Saccharomyces cerevisiae.
The yeast Scheffersomyces stipitis, however, is well known for its high native capacity to ferment xylose. Here, researchers at the Chinese Academy of Agricultural Sciences applied next-generation sequencing technology for RNA (RNA-Seq) to generate two high-resolution transcriptional maps of the S. stipitis genome when this yeast was grown using glucose or xylose as the sole carbon source. RNA-Seq revealed that 5,176 of 5,816 annotated open reading frames had a uniform transcription and that 214 open reading frames were differentially transcribed. RNA-Seq Blog Poll Results. Seq used to Trim the Mollusc Family Tree. Molluscs (snails, octopuses, clams and their relatives) have a great disparity of body plans and, among the animals, only arthropods surpass them in species number.
This diversity has made Mollusca one of the best-studied groups of animals, yet their evolutionary relationships remain poorly resolved. Open questions have important implications for the origin of Mollusca and for morphological evolution within the group and attempts to understand the early evolution of molluscs become even more complex when considering the large diversity of Cambrian fossils. To better resolve the relationships among molluscs, a group led by researchers at Brown University used RNA-Seq to generate transcriptome data for 15 species that, in combination with existing data, represent for the first time all major molluscan groups. This well-resolved tree will constitute a framework for further studies of mollusc evolution, development and anatomy. Alternative Splicing – RNA-Seq. RNA-Seq of the yeast Scheffersomyces stipitis. Seq Workshop – An introductory course to RNA-Seq. Video Protocol – Single Read and Paired End mRNA-Seq Illumina Libraries from 10 Nanograms Total RNA.
The authors describe a protocol for the Illumina Genome Analyzer II platform for mRNA-Seq sequencing for library preparation that avoids significant PCR amplification and requires only 10 nanograms of total RNA.
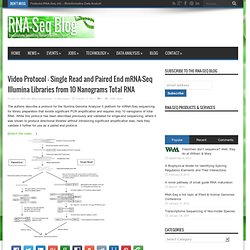
While this protocol has been described previously and validated for single-end sequencing, where it was shown to produce directional libraries without introducing significant amplification bias, here they validate it further for use as a paired end protocol. (Watch the video… ) Video Protocol – Single Read and Paired End mRNA-Seq Illumina Libraries from 10 Nanograms Total RNA.
Another weapon in the war against bias. There have been several studies demonstrating the biases inherent to the RNA-Seq method as well as variation in results across protocols and platforms. Researchers have set about innovating methods to correct for these biases and variances, but until now, most correction methods involve the use of bioinformatics models for partial correction. (See Post – Bias Detection and Correction in RNA-Sequencing Data) Flowcell Reverse Transcription Sequencing (FRT-seq) PCR is an inherently biased process, which decreases the efficiency of transcriptome sequencing data acquisition. Flowcell reverse transcription sequencing is a method of transcriptome sequencing for Illumina sequencers in which the reverse transcription reaction is performed on the flowcell by using unamplified, adapter-ligated mRNA as a template. This approach removes PCR biases and duplicates, generates strand-specific paired-end data and is highly reproducible. The procedure can be performed quickly, taking 2 d to generate clusters from mRNA.
Mamanova L, Turner DJ. (2011) Low-bias, strand-specific transcriptome Illumina sequencing by on-flowcell reverse transcription (FRT-seq). RNA CaptureSeq for targeted RNA sequencing of the human transcriptome. Seq greatly improves the accuracy of prediction of protein-coding genes. As more and more genomes are sequenced, genome annotation becomes increasingly important in bridging the gap between sequence and biology. Gene prediction, which is at the center of genome annotation, usually integrates various resources to compute consensus gene structures. However, many newly sequenced genomes have limited resources for gene predictions. A group led by researchers at Beijing Normal University set out to create high-quality gene models of the cucumber genome (Cucumis sativus var. sativus), based on the EVidenceModeler gene prediction pipeline.
Using RNA-Seq reads of 10 cucumber tissues, they were able to reassemble the cucumber genome. They applied the new pipeline to the reassembled cucumber genome and included a comparison between their predicted protein-coding gene sets and a published set. RNA-Seq Blog Poll Results. Seq used to Trim the Mollusc Family Tree. The Transcriptome of the Potato – Solanum tuberosum. Seq greatly improves the accuracy of prediction of protein-coding genes.
Transcriptome Sequencing Reveals Two Gene Families Prone to Rearrangement in Breast Cancer. RNA-Seq Satellite Workshop at ABRF 2012. RNA-Seq – Insights into the Active Genome. Integrated RNA-Seq Data Analysis Pipeline – Generating Evidence-Backed Hypotheses in Silico. New RNA-Seq Chapters – Methods in Molecular Biology. Seq: A practical guide to the analysis of differential gene expression. Alternative Splicing – RNA-Seq. Transcriptome Sequencing of the Oilseed Rape. Advanced RNA-Seq and ChiP-Seq Data Analysis – Hands-on training at EBI. New RNA-Seq Chapters – Methods in Molecular Biology. Human Transcriptome Array Outperforms RNA-Seq. Comprehensive RNA-Seq Data Presentation with Comparison to Microarrays. RNA-PET – Genome Wide Full-Length Transcript Analysis Using 5′ and 3′ Paired-End-Tag RNA-Seq. A reference transcriptome for the cauliflower coral – Pocillopora damicornis.
New RNA-Seq template preparation protocol enables detection of imprinted macro ncRNAs. New RNA-Seq template preparation protocol enables detection of imprinted macro ncRNAs. ReCount – A multi-experiment resource of analysis-ready RNA-seq gene count datasets. RNA CaptureSeq for targeted RNA sequencing of the human transcriptome. Introduction to transcriptome analysis using HighThroughput Sequencing technologies (HTS) Will genome sequencing be routine in cancer care and research? Seq Hands-on Workshop. Genome sequence of an Australian kangaroo, Macropus eugenii, provides insight into the evolution of mammalian reproduction and development.