

GiantVirus. Genome Sequence of Mimivirus. The Discovery and Characterization of Mimivirus, the Largest Known Virus and Putative Pneumonia Agent. + Author Affiliations Reprints or correspondence: Dr.
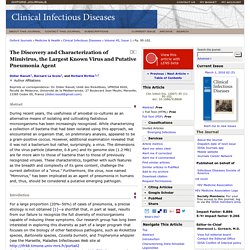

Didier Raoult, Unité des Rickettsies, UPRESA 6020, Faculté de Médecine, Université de la Méditerrannée, 27 Boulevard Jean Moulin, Marseille, 13385 Cedex 05, France (didier.raoult@gmail.com) Abstract During recent years, the usefulness of amoebal co-cultures as an alternative means of isolating and cultivating fastidious microorganisms has been increasingly recognized. While characterizing a collection of bacteria that had been isolated using this approach, we encountered an organism that, on preliminary analysis, appeared to be a gram-positive coccus. Introduction For a large proportion (20%–50%) of cases of pneumonia, a precise etiology is not obtained [1]—a shortfall that, in part at least, results from our failure to recognize the full diversity of microorganisms capable of inducing these symptoms. Figure 1 Strict intra-amoeba–growing microorganisms, as observed under light microscopy after Gram staining. mRNA deep sequencing reveals 75 new genes and a complex transcriptional landscape in Mimivirus.
+ Author Affiliations Abstract Mimivirus, a virus infecting Acanthamoeba, is the prototype of the Mimiviridae, the latest addition to the nucleocytoplasmic large DNA viruses.

The Mimivirus genome encodes close to 1000 proteins, many of them never before encountered in a virus, such as four amino-acyl tRNA synthetases. To explore the physiology of this exceptional virus and identify the genes involved in the building of its characteristic intracytoplasmic “virion factory,” we coupled electron microscopy observations with the massively parallel pyrosequencing of the polyadenylated RNA fractions of Acanthamoeba castellanii cells at various time post-infection.
Previous Gene Characterizations. Mimivirus in Pneumonia Patients - Volume 11, Number 3—March 2005. Giant mimi. Mimivirus gene promoters exhibit an unprecedented conservation among all eukaryotes. Proceedings of the National Academy of Sciences of the United States of Americawww.pnas.org Author Affiliations Communicated by James L.

Van Etten, University of Nebraska, Lincoln, NE, August 22, 2005 (received for review May 12, 2005) Abstract The initial analysis of the recently sequenced genome of Acanthamoeba polyphaga Mimivirus, the largest known double-stranded DNA virus, predicted a proteome of size and complexity more akin to small parasitic bacteria than to other nucleocytoplasmic large DNA viruses and identified numerous functions never before described in a virus.
Footnotes ↵ ‡ To whom correspondence should be addressed. Gene and Genome Duplication in Acanthamoeba polyphaga Mimivirus. Karsten Suhre* + Author Affiliations Gene duplication is key to molecular evolution in all three domains of life and may be the first step in the emergence of new gene function.

It is a well-recognized feature in large DNA viruses but has not been studied extensively in the largest known virus to date, the recently discovered Acanthamoeba polyphaga Mimivirus. Here, I present a systematic analysis of gene and genome duplication events in the mimivirus genome. I found that one-third of the mimivirus genes are related to at least one other gene in the mimivirus genome, either through a large segmental genome duplication event that occurred in the more remote past or through more recent gene duplication events, which often occur in tandem.
It has long been realized that new gene material frequently emerges through gene and genome duplication (25, 26). Detection of paralogous genes was performed using programs from the BLAST package (1). Received 27 June 2005. Mimivirus Giant Particles Incorporate a Large Fraction of Anonymous and Unique Gene Products. Ultrastructural Characterization of the Giant Volcano-like Virus Factory of Acanthamoeba polyphaga Mimivirus.
Abstract Acanthamoeba polyphaga Mimivirus is a giant double-stranded DNA virus defining a new genus, the Mimiviridae, among the Nucleo-Cytoplasmic Large DNA Viruses (NCLDV).
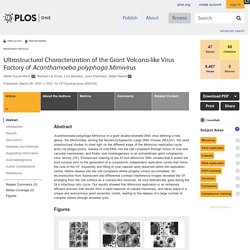
We used utrastructural studies to shed light on the different steps of the Mimivirus replication cycle: entry via phagocytosis, release of viral DNA into the cell cytoplasm through fusion of viral and vacuolar membranes, and finally viral morphogenesis in an extraordinary giant cytoplasmic virus factory (VF). Fluorescent staining of the AT-rich Mimivirus DNA showed that it enters the host nucleus prior to the generation of a cytoplasmic independent replication centre that forms the core of the VF.
Assembly and filling of viral capsids were observed within the replication centre, before release into the cell cytoplasm where progeny virions accumulated. 3D reconstruction from fluorescent and differential contrast interference images revealed the VF emerging from the cell surface as a volcano-like structure. Results A. Distinct DNA Exit and Packaging Portals in the Virus Acanthamoeba polyphaga mimivirus. PLOS Biology: Structural Studies of the Giant Mimivirus. Abstract Mimivirus is the largest known virus whose genome and physical size are comparable to some small bacteria, blurring the boundary between a virus and a cell.
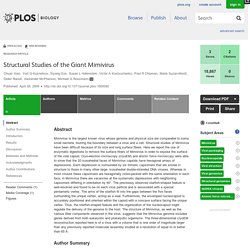
Structural studies of Mimivirus have been difficult because of its size and long surface fibers. Here we report the use of enzymatic digestions to remove the surface fibers of Mimivirus in order to expose the surface of the viral capsid. Cryo-electron microscopy (cryoEM) and atomic force microscopy were able to show that the 20 icosahedral faces of Mimivirus capsids have hexagonal arrays of depressions. Each depression is surrounded by six trimeric capsomers that are similar in structure to those in many other large, icosahedral double-stranded DNA viruses.
Author Summary Mimiviruses are larger than any other known virus, yet despite their size, the capsid has been shown to be a regular icosahedron. The Three-Dimensional Structure of Mimivirus. Morphogenesis of Mimivirus and Its Viral Factories: an Atomic Force Microscopy Study of Infected Cells. Mimivirus (Acanthamoeba polyphaga mimivirus) is one of nature's largest characterized viruses, having a diameter of 700 to 750 nm based on electron microscopy (EM) (1–3) and atomic force microscopy (AFM) (4), with a genome size of 1.2 million base pairs (5), nearly twice that of any other known virus.

Its linear, double-stranded DNA codes for 911 genes, and among these are many that are not expressed by any other virus but only by living cells. It is the focus of particular interest because its genomic, metabolic, and structural intricacies suggest that it might provide an evolutionary bridge between viruses and living cells (6–9). Mimivirus infects the amoeba Acanthamoeba polyphaga, although its host range is likely to be considerably larger (10). It appears to be most closely related structurally to large algal viruses such as Paramecium bursaria chlorella virus 1 (PBCV-1) and other iridoviruses (11–13), although it possesses additional features not present in those viruses. Fig 1.
Legionella.