

Www.watchmouse.com/en/SPI/2010/DAX30_spi.png. By Life Technologies. Guides. Ion Torrent. Ion Torrent. A first peek at data from an Ion 316 chip. Ion Torrent has made available an E.coli DH10B fragment dataset for the 316 chip, which is expected to be generally available early next month.
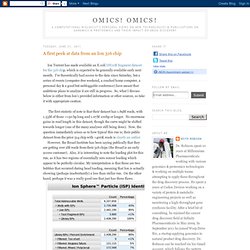
I've theoretically had access to the data since Saturday, but a series of events (computer-free weekend, a crashed home computer, a personal day & a good but unbloggable conference) have meant that ambitious plans to analyze it are still in progress. So, what I discuss below is either from Ion's provided information or other sources, so take it with appropriate caution. The first statistic of note is that their dataset has 1.69M reads, with 1.53M of those >=50 bp long and 1.07M 100bp or longer. No enormous gains in read length in this dataset, though the curve might be shifted towards longer (one of the many analyses still being done). Now, the question immediately arises as to how typical this run is; their public dataset from the prior 314 chip with >400K reads is clearly an outlier.
Ionjuly11releasefinal.pdf (application/pdf Object) Ion Community Home: Home. Ion OneTouch. Ion Community Home: Video: Ion Xpress Template Kit - hear what people are saying. Ion Community Home: Video: Ion Torrent Technology: How is it made? Ion Community Home: Video: Ion Torrent Technology: How does it work ? Poster30.ashx (application/pdf Object) Ion Community Home: Video: Ion Torrent Webinar - March 2011. Homopolymer tails. Homopolymer tails homopolymer tails a segment containing several of the same kind of deoxyribonucleotides arranged in tandem at the 3′ end of a DNA strand.
See Appen- dix C, 1972, Lobban and Kaiser; terminal transferase. Homo sapiens the species to which human beings belong. The scientific name was coined in 1758 by Linnaeus. The estimated genome size is 3.2 × 109 base pairs. Compare with isogamy. homothallic fungus a fungal species producing a sexual spore that results from the fusion of geneti- cally different nuclei derived from the same thallus. See Appendix C, 1900, Bateson. homozygous having identical rather than different alleles in the corresponding loci of homologous chromosomes and therefore breeding true. Malted barley is used in the brewing industry, and food-grade malts are components in the formulation of baked goods and breakfast cere- als. In prokaryotes, ex- amples of HMEs are F factors, R (resistance) plas- mids, and prophages (all of which see). Hox genes Random Posts. An integrated semiconductor device enabling non-optical genome sequencing : Nature.
IonTorrent: Benchtop Sequencing, Streamlined. New, smaller sequencing instruments are finding niches everywhere.

Earlier this year in Less Is More: Sequencing on the Benchtop, I profiled three pint-sized sequencing instruments designed for next-gen sequencing on the benchtop: IonTorrent’s PGM, the Illumina MiSeq, and 454′s GS Junior. I must admit that I’m most intrigued by the PGM, probably because the idea of semiconductor sequencing is just so fascinating. One concern that I’ve heard about the PGM, however, was sample preparation. The time requirement and difficulty of sample prep was an immediate question at AGBT when Jonathan Rothberg first presented the instrument. He dodged the question, and I don’t think there’s been a satisfactory answer since. Priced at under $5,000, and measuring 1′ x 1′ square, the OneTouch requires “five minutes of hands-on time” and takes a few hours to run. PGM Applications Essentially, the niche for PGM, MiSeq, and GS Junior is everything that’s not quite enough for a full-on sequencing run. First Look: Data from IonTorrent’s 316 Chip.
I got an early look at data from IonTorrent’s forthcoming semiconductor sequencing chip, called the 316.
The data comes from a single sequencing run of E. coli, specifically the laboratory reference strain DH10B, not to be confused with the hybrid E. coli strain in the European outbreak that IonTorrent sequenced earlier this year. According to the run report, some 1.69 million reads were generated, of which 1.35 million were 100 base pairs or longer. It’s a nice, shiny report with color images, and of course I summarily ignored it in favor of looking at the data myself.
The data package included an SFF file, a data format often used for Roche/454 data that contains read sequences and base qualities. There was also a BAM file, which included the read sequences aligned to the E. coli reference, indexed for random access. Life Technologies. Job als Account Manager / Sales Rep Ion Torrent Germany » Experteer.
Diese Stelle ist nicht mehr verfügbar!
Aktuelle Jobs. Escherichia coli O104:H4 str. TY-2482, whole genom... - Nucleotide result. Prospective Genomic Characterization of the German Enterohemorrhagic Escherichia coli O104:H4 Outbreak by Rapid Next Generation Sequencing Technology. An ongoing outbreak of exceptionally virulent Shiga toxin (Stx)-producing Escherichia coli O104:H4 centered in Germany, has caused over 830 cases of hemolytic uremic syndrome (HUS) and 46 deaths since May 2011.

Serotype O104:H4, which has not been detected in animals, has rarely been associated with HUS in the past. To prospectively elucidate the unique characteristics of this strain in the early stages of this outbreak, we applied whole genome sequencing on the Life Technologies Ion Torrent PGM™ sequencer and Optical Mapping to characterize one outbreak isolate (LB226692) and a historic O104:H4 HUS isolate from 2001 (01-09591). Reference guided draft assemblies of both strains were completed with the newly introduced PGM™ within 62 hours. Open-Source Genomic Analysis of Shiga-Toxin–Producing E. coli O104:H4.
Nutzerprofil: Keith Robison. Education Home. Nature.