

GeneDesigner2. The DNA design tool for today’s molecular biologists Capture your entire gene design process in one efficient application, using a range of design tools purposely built for the task: Intelligent, fast, and easy-to-use algorithms for in silico cloning, codon optimization, back translation and primer design.
Graphically rich molecular view to display, annotate and edit your constructs. Customizable database to quickly store, manage, and track genetic element, genes and constructs. Powerful, intuitive drag-and-drop interface for moving sequence elements within or between constructs. Gene Designer Sequence View enables easy manipulation of sequence elements, codon choices, and oligonucleotide positions.
You Dream It. Assembling genetic building blocks using Gene Designer 2.0 enables you to design, clone and validate in powerful new ways. Codon optimize for recombinant protein expression in any organism using multiple codon-optimization algorithms. The DNA design tool for tomorrow’s scientists. Trends in Cell Biology - Table of Contents - Volume 22 Issue 12, December 2012. DOI: DOI: xTypically, findings from cell biology have been beneficial for preventing human disease.
However, translational applications from cell biology can also be applied to conservation efforts, such as protecting coral reefs. Recent efforts to understand the cell biological mechanisms maintaining coral health such as innate immunity and acclimatization have prompted new developments in conservation. Similar to biomedicine, we urge that future efforts should focus on better frameworks for biomarker development to protect coral reefs.
DOI: xA major component of cellular aging is the detrimental accumulation of mutations in the cell's DNA. Welcome to the mcmanus lab! Table of Contents — July 2012, 40 (W1) InterPreTS - Interaction Prediction through Tertiary Structure. IBIS. BBCU - Sequence Analysis/Promoters. Remotely Activated Protein-Producing Nanoparticles - Nano Letters. Avi Schroeder †‡, Michael S.
Goldberg †, Christian Kastrup †‡§, Yingxia Wang ‡, Shan Jiang †‡, Brian J. Joseph ‡, Christopher G. Levins †‡, Sneha T. Kannan †, Robert Langer †‡ , and Daniel G. †David H. . § Michael Smith Laboratories and Department of Biochemistry and Molecular Biology, University of British Columbia, Vancouver V6T 1Z4 Canada Harvard MIT Division of Health Science and Technology, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, United States Nano Lett., 2012, 12 (6), pp 2685–2689 DOI: 10.1021/nl2036047 Publication Date (Web): March 20, 2012 Copyright © 2012 American Chemical Society Author Contributions These authors contributed equally to this work.
Section: Abstract The development of responsive nanomaterials, nanoscale systems that actively respond to stimuli, is one general goal of nanotechnology. Citing Articles View all 15 citing articles Citation data is made available by participants in CrossRef's Cited-by Linking service. Solve for X: Juan Enriquez on harnessing synthetic genetics. GeNorm normalization of real-time PCR expression data. [introduction] geNorm is a popular algorithm to determine the most stable reference (housekeeping) genes from a set of tested candidate reference genes in a given sample panel.
From this, a gene expression normalization factor can be calculated for each sample based on the geometric mean of a user-defined number of reference genes. The Microsoft Excel geNorm version from 2002 has been downloaded more than 15,000 times worldwide. Since 2010, a much improved geNorm module is integrated in the qbase+ software (both basic and premium license) for real-time PCR data analysis in Windows, Mac and Linux (available from Biogazelle).
The manual is available here (requires free login to the MyBiogazelle community). The underlying principles and formulas are described in Vandesompele et al., Genome Biology, 2002, 'Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes'. [geNorm citations] [Improved geNorm in qbase+ software] 1. [feedback] Entenda a Engenharia Metabólica. Uma das grandes maravilhas da humanidade – objeto de grande satisfação entre os químicos – é uma tabela que nos diz tudo o que existe no universo, os cerca de 120 elementos que formam tudo aquilo que o ser humano conseguiu perceber.
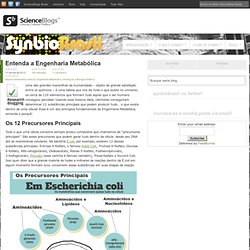
Usando essa mesma ideia, cientistas conseguiram determinar 12 substâncias principais que podem produzir tudo… o que existe dentro de uma célula! Esse é um dos princípios fundamentais da Engenharia Metabólica, entenda o porquê: Os 12 Precursores Principais Tudo o que uma célula consome sempre produz compostos que chamamos de “precursores principais”. São esses precursores que podem gerar tudo dentro da célula: desde seu DNA até às membranas celulares. Assim, ao melhor estilo dos antigos alquimistas, pesquisadores – em especial FC Neidhardt – dissecaram células de E.coli de modo a determinar a quantidade desses precursores que seria necessária para “construir” uma bactéria (ver infográfico acima): O Fluxoma Arte de Pedro Pantai.
Fluxos Metabólicos Referências. PathwayAccess. Everybody is using one of many different pathway databases.

We need a tool to integrate pathways from different sources. Please cite: PathwayAccess: CellDesigner Plugins for Pathway Databases John L. Van Hemert; Julie A. Dickerson Bioinformatics 2010; doi: 10.1093/bioinformatics/btq423 NEW! Introduction PathwayAccess is a suite of CellDesigner plugins which directly interact with pathway datasources so that the user can accomplish one or both of the following: